Gene isolation and characterisation, gene expression, gene regulation in plants
Theory
In order to function, genes have to be expressed. The process by which the instructions contained inside a gene are utilized to produce gene products, which are primarily proteins but can also include functional RNA molecules, is referred to as gene expression. This process is essential to the operation of cells and organisms because it determines the proteins that are made and, as a result, the functions that a cell is able to carry out.
Importance of Gene expression –
- The protein that a cell makes determines its shape and function. These proteins are identified with the help of gene expression. Different gene expression patterns allow cells to specialize and differentiate, giving rise to different tissues and organs with unique roles.
- Gene expression enables cells to react to changes in their environment, including the availability of nutrients, stress, and messages from other cells. The ability to adapt is essential for both survival and homeostasis.
- Misregulation of gene expression can lead to diseases, including cancer, genetic disorders, and metabolic conditions. Understanding gene expression patterns helps in diagnosing, treating, and developing therapies for these diseases.
- In order to control metabolic pathways and preserve homeostasis, gene expression is essential. Gene expression produces regulatory proteins, hormones, and enzymes that regulate several physiological processes.
- The majority of the activities carried out by cells, such as initiating metabolic reactions, replicating DNA, reacting to stimuli, and moving molecules, are directly attributed to the process of gene expression.
Different types of Genes & Gene Expression, for better understanding –
Constitutive Gene
Constitutive genes encode essential cellular functions and are thus, always expressed continuously and are always active.
EXAMPLE – Actin, particularly β-actin, along with glyceraldehyde-3-phosphate dehydrogenase (GAPDH), tubulin, cyclophilin, elongation factor 1-α (ef1α), ubiquitin, and 18 Svedberg Units (S) rRNA (18S rRNA), are constitutively expressed and participate in fundamental housekeeping functions essential for cellular maintenance.
Tubulin genes in plants help in control of cell shape and intracellular support.
ILLUSTRATION - Constitutive genes are like the utilities (electricity, water) in a house that are always on to keep the house running smoothly.Inducible Gene
These are expressed only under specific conditions, they are turned on in response to an inducer molecule or environmental stimulus.
EXAMPLE – AlcR/AlcA (ethanol inducible); GR fusions, GVG, and pOp/LhGR (dexamethasone inducible); XVE/OlexA (β-estradiol inducible); and heat shock induction.
ILLUSTRATION -Inducible genes are like a security system that activates when a burglar tries to break in, alerting the occupants to take action.Repressible Gene
Repressible genes are normally active, but in the presence of a certain chemical, they can become inactive, or repressed. These genes frequently function in pathways that are anabolic.
EXAMPLE – Nitrate Reductase Genes – these are generally involved in nitrogen metabolism and are repressed when nitrogen levels are high, Nac 2 gene
ILLUSTRATION – Repressible genes are like a factory that shuts down when the warehouse is full.Tissue Specific Gene
Tissue-specific genes are expressed only in particular tissues, allowing for the specialization of cells within different parts of the body.
EXAMPLE – Storage protein Genes in Seeds – these are expressed specifically in seeds and encode proteins that store nutrients. PHA (Phaseolin) in beans is one such example, EMB-23 gene in ripe pulp and flower of Grand Naine(Banana), Petal development in orchids is governed by high expression of clade-1/2 DEF like genes and low expression of clade-3/4 DEF-like genes. The transitory expression of the OgCHS gene from Gower Ramsey in the white Honey Dolls of the Oncidium species is responsible for the production of Anthocyanins which display red patterns. Expression of ethylene insensitive-3 like (EIL) and ethylene response factor (ERF) is responsible for fruit ripening in Peach.Developmentally Regulated Gene
There are some genes that are expressed at specific stages of organism’s development. They further ensure proper progression of developmental stages.
EXAMPLE – Leafy (LFY) Gene – these are critical for the transition from vegetative to floral development in plants, Baby Boom (BBM) gene regulates early embryo and endosperm development, Transparent testa glabra2 (TTG2) controls trichome development and seed coat production, KNAT1 induces lobed leaves with ectopic meristems when overexpressed in Arabidopsis.
Gene expression process
Gene expression process consists of – transcription and translation. Specific sections of the genome known as "transcription units" act as templates for the creation of ribonucleic acid (RNA), a different type of nucleic acid.
There are 3 types of RNA –
- rRNA (Ribosomal RNA) – structural component of ribosomes.
- tRNA (Transfer RNA) – involved in protein synthesis (translation).
- mRNA (Messenger RNA) – carries the protein-coding information from DNA and is translated subsequently for protein synthesis.
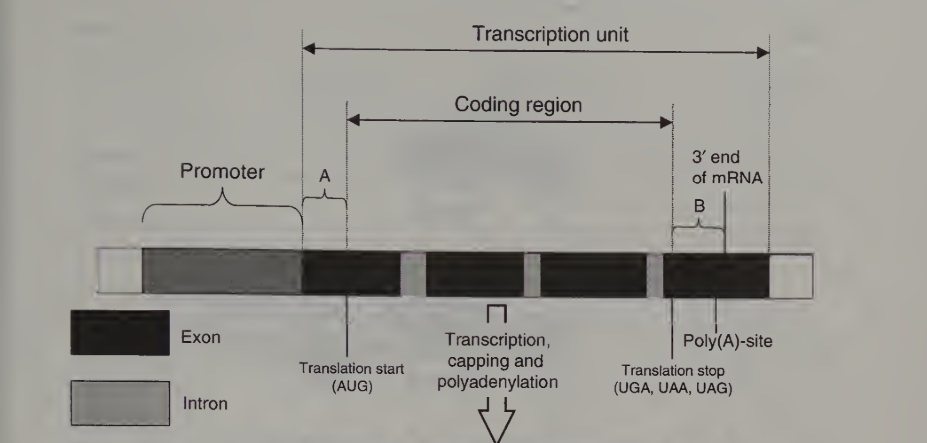
Figure 1. The structure and expression of a eukaryotic protein-coding gene
Structure of the protein coding gene
Since, the genes that encode proteins are largely dealt with, it is crucial to understand the structure of the protein coding gene.
Each transcription region is associated with a regulatory region called as the promoter (flanking region at the 5’ end of a gene). Promoter is important as it contains the binding site for RNA polymerase II enzyme which is responsible for making RNA copy of the DNA during “transcription”.
The transcription unit region of the gene is transcribed by the above enzyme into pre – mRNA. This pre – mRNA region consists of – 5’ UTR (A in the figure), the coding region and a 3’ UTR (B).
The coding region is further composed of extrons and introns. Extrons carry information related to the protein sysnthesis, it is called as “coding” as it is translated. Introns on the other hand are “non-coding”.
The 3’ end of the gene contains a poly (A) site (AATAAA) which acts as a signal for the adenylation of the RNA. During the processing of pre –mRNA, the RNA is cleaved and is also modified by addition of a ‘cap’ of 7- methylguanosine linked to pre –mRNA by three phosphate group.
Further, splicing takes place where the introns are removed and extrons are joined together.
Translation
In this process the nucleotide ‘language’ of DNA and RNA is converted into amino acid language of proteins.
The processed mRNA is transported to the ribosome, where the ribosome's small subunit binds to the mRNA. The initiator tRNA, carrying methionine, binds to the start codon (AUG) on the mRNA. Further, the ribosome moves along the mRNA, adding amino acids one by one to the growing polypeptide chain. Each amino acid is brought to the ribosome by a specific tRNA molecule, which pairs with the corresponding codon on the mRNA. When the ribosome reaches a stop codon (UAA, UAG, or UGA), translation ends. The newly synthesized polypeptide is released, and the ribosomal subunits dissociate from the mRNA. Numerous mechanisms, such as chromatin structure, transcriptional control, RNA processing, RNA stability, translational control, and post-translational modifications, are involved in the regulation of gene expression. The proper genes are expressed at the appropriate time and in the appropriate quantity due to this regulation, which enables cells to react to both internal and external cues.
How gene expression is critical for plant growth and development?
The majority genes of chloroplast genomes are crucial for plant survival as they encode essential subunits of the photosynthetic apparatus and the chloroplast gene expression system, including RNA polymerase subunits, ribosomal proteins, ribosomal RNAs (rRNAs), and transfer RNAs (tRNAs). Significant modification of chloroplast gene expression can be lethal to plants, highlighting the critical role of these genes.
Similarly, NFL (No- flowering in short day) is a transcription factor that promotes flowering in Arabidopsis under short-day conditions. In the absence of NFL, plants do not flower under short-day conditions.
MicroRNAs regulate gene expression networks essential for appropriate development, cellular survival, and stress responses. miRNAs can engage with hormones to assist plants in regulating their growth, development, and differentiation.
