Gene isolation and characterisation, gene expression, gene regulation in plants
Overview
What is a Gene?
Gene is a unit of inheritance. It can be defined as a segment of the genome with a specific sequence of nucleotides, with a specific biological function.
The genome of any living organism is its genetic foundation, consisting of a lengthy sequence of deoxyribonucleic acid (DNA). This DNA contains the entirety of the organism's hereditary information, which is passed down to its individual cells. The genome consists of chromosomal DNA, plasmid DNA, and organelle DNA (in eukaryotes), found in mitochondria and chloroplasts.
The growth and development of the organism is determined by the products resulting from the expression of nucleotide sequences inside the genome. The DNA sequence orchestrates the synthesis of all the ribonucleic acids (RNAs) and proteins in the organism, ensuring that they are produced in the correct cells and at the right time through an intricate set of connections.
Genes are responsible for the production of RNA. When this occurs, the RNA is then utilised to control the creation of a polypeptide in many instances. In other instances, such as transcription of ribosomal RNA (rRNA) and transfer RNA (tRNA) genes, the RNA that is produced as a result of the gene's transcription is the functional end product. A sequence of DNA that encodes an RNA is referred to as a gene. In the case of structural genes, which are responsible for encoding proteins, the RNA in turn encodes a polypeptide chain.
Do you Know?
- If the entire pea DNA were expressed, it would represent 4 million genes.
- Plants possessing a compact genome and a little amount of repetitive DNA exhibit brief somatic and meiotic cycles, as well as a quick generation time. An example here is Arabidopsis thaliana, which is widely utilised in the field of plant molecular biology due to its low DNA content and short generation period of five weeks
- Rice has a relatively small genome size of 415 Mbp, in contrast to wheat which has a much larger genome size of 16 billion base pairs.
Gene Structure
Plant genes have a complicated structure made up of several essential elements that are involved in the transcription, translation, and control of genetic information. Transcription of genes results in mRNA, which is then translated into polypeptides. In case of ribosomal RNA (rRNA) and transfer RNA (tRNA) there is transcription, but they are not translated into polypeptides. These non-coding RNAs carry out a variety of tasks for the cell.
- Promoter region
The promoter is a distinct DNA sequence situated upstream (towards the 5' end) of the gene's coding region. The binding of RNA polymerase to these promoters is crucial for the commencement of transcription. The transcription rate of a specific DNA segment is determined by the sequence of bases in its promoter. The DNA sequence at a promoter site differs among genes, although all promoters possess a consensus sequence.
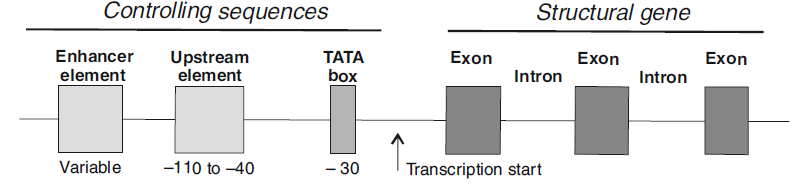
Figure 1. Diagrammatic representation of a typical nuclear gene
TATA box
The TATA-box, also known as the Goldberg Hogness box, is a short sequence element that is commonly found upstream from the cap site in many eukaryotic genes. It plays a crucial role in proper transcription. Among these sequences, the TATA box is the most extensively studied and understood.
In nearly all genes from plants and animals, this region typically occurs -25 to -30 nucleotides upstream of the cap site. This series is necessary for both in vitro and in vivo accurate expression of eukaryotic genes.
(The negative numbers indicate the relative location of each box with respect to the starting point of transcription, which is indicated as +1.The +1 position refers to the place where the first nucleotide is added to the encoded RNA molecule).
AGGA box
The consensus sequence "CAAT" box is potentially involved in regulating the transcription of several eukaryotic genes. The sequence is GG (CT) CAATCT. This sequence is commonly located at positions -40 and -180 nucleotides upstream of the cap site. The arrangement of these pieces is quite diverse, however it frequently consists of the sequence CAAT, which is referred to as the CAAT box. In the context of plants, the AGGA box is the term used to describe it.
Enhancer Sequence
Enhancer sequences are a distinct category of elements that exhibit a significant amount of transcription.
The origins of these sequences were initially discovered in viral genes but they have also been discovered in plants. Enhancers are situated within a short distance of a few hundred nucleotides before the initiation of transcription. While enhancer elements are typically located in the flanking sequences at the 5′ ends of genes, they can also be found in other locations.
A specific enhancer core sequence, such as GGTGTGGAAAG, or more generally GTGGT/AT/AT/AG (written as GTGGWWWG, where T/A is represented as W), can be identified through sequence comparisons. However, it is evident that other sequences besides the core are also significant. Plant enhancer-like sequences promote the particular expression of genes in tissues.
- 5’ UTR region
The 5' untranslated region (5' UTR) is an essential component of the gene that performs multiple crucial functions in the control of gene expression. The 5' UTR, despite not being translated into protein, plays a crucial role in regulating the efficiency and precision of translation. The 5' untranslated region (UTR) is positioned between the promoter region and the initiation codon (AUG) of the gene.
The main purpose of the 5' UTR is to control the start of translation. This regulation can be achieved through various mechanisms:
Ribosome binding and scanning involve the ribosome scanning the 5' untranslated region (UTR) in order to locate the initiation codon. The existence of secondary structures or regulatory elements can impact the ribosome's ability to locate the start codon.
Translational repression occurs when certain sequences or structures in the 5' UTR region of mRNA bind to regulatory proteins or micro RNAs, resulting in the inhibition of translation.
- Coding Sequences
Exons are the portions of a gene that carry biological information or coding sequences, and the intervening sequences are referred to as Introns.
These discontinuous genes are also known as split or mosaic genes, and are now recognised to be prevalent in higher species. These introns are transcribed, but they are not translated since they are not seen in mature mRNA. While they have been shown in some plant structural genes, introns have not been discovered in rRNA genes. Introns can range from zero in higher species to hundreds in some. Introns are frequently far longer than exons.
Using genomic clones to compare the nucleotide sequences of DNA and mRNA, introns can be found. Under an electron microscope, cloned genomic DNA is annealed with RNA and the resulting material is analysed. The intron sequences will create single-stranded loops in the heteroduplex molecule since they are not present in the RNA.
There is a wide range in the number of introns.
Many plant genes, such as the soybean protein genes, the Zein storage genes of maize, the uncommon class leaf gene, and the majority of cab genes (nuclear genes generating the photosystem's chlorophyll a/b binding protein) have no introns at all. Soybean actin, soybean leghemoglobin, soybean glycinine, a rare class leaf gene, and the small subunit of RuBP carboxylase are examples of plant genes containing two or three introns. The phytochrome gene has five introns, while the maize Adh gene has nine.
All plant genes are the same in the first and last two bases of the introns GT and AG, respectively, when intervening sequences are present. All eukaryotic genes share the same two dinucleotides at the intron/exon junctions, indicating that comparable RNA splicing mechanisms are at work.
3’ UTR
The 3' UTR is not translated into protein and comes after the stop codons (UAA, UAG, and UGA). This area is home to sequences that both indicate the end of transcription and are important for nuclear export, translation efficiency, and mRNA stability.
(Stop codon & Start Codon - 61 of the 64 potential codons designate a specific amino acid; the other three codons (UAA, UAG, and UGA) serve as stop signals, alerting the translation machinery to the completion of protein synthesis.
Since the codon AUG in the mRNA determines the start site for translation in nearly all circumstances, the codon AUG (which specifies methionine) is also referred to as a start codon. It should be noted that all other amino acids have several codons, while only two—tryptophan and methionine—have just one. )Polyadenylation Signal
This sequence is situated in the 3' untranslated region (UTR) and serves as a signal for the attachment of a poly(A) tail to the mRNA transcript. The presence of a poly(A) tail is essential for maintaining the stability of mRNA, facilitating its export from the nucleus, and enabling the process of translation.Termination site
This signifies the conclusion of the transcription process. The process entails distinct sequences that prompt RNA polymerase II to halt transcription and liberate the recently produced mRNA.
