Metabolic flux determination, pathway engineering in plants
Theory
Metabolic Flux Analysis (MFA) in Plants
Metabolic flux refers to the rate at which metabolites (the intermediates and products of metabolism) are converted through a metabolic pathway within a biological system. It is a quantitative measure of the flow of biochemical substances through metabolic networks, reflecting the dynamic state of cellular metabolism. Metabolic flux is expressed in terms of concentration per unit time (e.g., mmol/gDW/h, where gDW = grams of dry weight of cells per hour). It provides a rate, not just a static snapshot, indicating how active a particular pathway or reaction is under specific conditions. It is defined for individual reactions or entire pathways within the metabolic network. Metabolic fluxes are influenced by nutrient availability, genetic regulation, enzyme activity, energy status, and environmental factors (e.g., oxygen levels, pH). In systems biology, fluxes help model and understand the overall behaviour of metabolic networks. In metabolic engineering, controlling or rerouting fluxes is key to optimizing production of desired metabolites (e.g., pharmaceuticals, biofuels).
Let us consider an example of riding a bicycle. The goal here is to move forward. There are multiple control points in a bicycle, which controls its overall function, the forward motion – the pedals, the handle and the brakes. The pedals are controlled by the mechanical force applied by the human legs – a higher force on the pedals results in a faster forward motion. This could be imagined as an overexpression of a gene coding for an enzyme responsible for catalysing a rate limiting step in a metabolic pathway, resulting in boosting the rate of reaction. On the other hand, the brakes are applied to slow down the forward motion of the bicycle when required, which can be imagined as a gene silencing event, which slows down the reaction rate. All these control points are coordinated in such a way that the forward motion of the bicycle is present in a desired way depending upon the circumstances. Same is the case for any metabolic reaction is a biological system.
Metabolic Flux Analysis (MFA) in plants is a quantitative method employed to assess the rates of metabolite movement via diverse metabolic pathways, offering a dynamic representation of cellular metabolism. In contrast to static metabolomics, which assesses metabolite concentrations at a singular time point, MFA records the real-time activity and directionality of metabolic events, providing enhanced understanding of pathway operation and regulation. This is especially significant in plants, where metabolism is extensively compartmentalized across organelles including chloroplasts, mitochondria, and peroxisomes. A fundamental element of MFA is the use of stable isotope-labelled substrates, such as ^13C-glucose or ^13CO₂, administered to plant tissues or cells; the ensuing distribution of these isotopes in metabolic intermediates is subsequently examined by mass spectrometry or NMR. By amalgamating these labelling patterns with computational modelling, researchers may rebuild flux maps that elucidate bottlenecks, alternate pathway utilization, and metabolic efficiency. MFA has been instrumental in unravelling how plants allocate carbon and energy under different environmental conditions, how they respond to stress, and how they can be engineered for improved productivity or synthesis of valuable compounds. Nonetheless, obstacles include spatial complexity, tissue heterogeneity, and technical constraints in delineating compartment-specific flows persistently stimulate innovation in this domain.
The notion of metabolic flux—the transit of metabolites through a biochemical pathway—was challenging to examine in real-time prior to the 1940s. The introduction of radioisotopes enabled researchers to effectively track the movement of atoms inside metabolic networks. Initial research utilizing isotopes such as carbon-14 (¹⁴C), phosphorus-32 (³²P), and oxygen-18 (¹⁸O) provided unparalleled understanding of metabolic pathways. This method was revolutionary because It offered temporal resolution, enabling researchers to monitor metabolites in real-time over seconds, it provided spatial specificity: atoms from labelled substrates could be tracked into particular molecules and facilitated quantitative assessments of metabolic rates (fluxes), rather than only static concentrations.
One of the major breakthroughs in the late 1940s and early 1950s was the discovery of the Calvin cycle - which was the first mapping of a carbon fixation pathway. Melvin Calvin, Andrew Benson, and James Bassham at the Radiation Laboratory, UC Berkeley discovered the Calvin cycle. Chlorella pyrenoidosa, a unicellular green alga was ideal for this experiment due to its rapid growth and single-cell nature. Carbon C14 was the radioisotope used - carbon-14 labelled CO₂ (¹⁴CO₂). The algae were subjected to ¹⁴CO₂ under light for brief durations, as little as 5 seconds. At each time point, cells were swiftly terminated using heated alcohol to "quench" metabolism, therefore effectively halting the current metabolic state. The metabolites were extracted and isolated via two-dimensional paper chromatography. The locations of ¹⁴C-labeled metabolites were depicted by autoradiography. The radioactivity at each location was measured to ascertain which chemicals were accumulating the label over time. The initial product to get a label was 3-phosphoglycerate (3-PGA), a molecule consisting of three carbon atoms. Labelling subsequently emerged in ribulose bisphosphate (RuBP), glyceraldehyde-3-phosphate (G3P), and fructose-6-phosphate, among others. From this temporal data, Calvin and his team inferred the cyclical process of carbon fixation: RuBP associates with CO₂, yielding 3-PGA, which is then decreased and ultimately regenerates RuBP. This piece of work is referred to as the Calvin–Benson–Bassham (CBB) cycle and constitutes the inaugural comprehensive pathway of carbon fixation ever documented. Melvin Calvin was awarded the 1961 Nobel Prize in Chemistry.
Notwithstanding the arduous and perilous characteristics of early radioisotope research, these endeavours delineated the comprehensive carbon fixation pathway, encompassing CO₂ absorption to sugar production. Identified the mechanism of oxygen evolution, validating water as the source. Disclosed the photosystems, electron transport chain, and energy conversion processes. Established the foundation for systems-level investigation of metabolism, since isotope tracing serves as a forerunner to contemporary metabolic flux analysis (MFA).
The initial investigations employing radioisotopes were technically challenging and necessitated both inventiveness and accuracy. Nonetheless, they substantially altered our comprehension of plant metabolism. These advancements persist in influencing contemporary study fields, ranging from synthetic biology to climate science, highlighting the enduring significance of these initial discoveries.
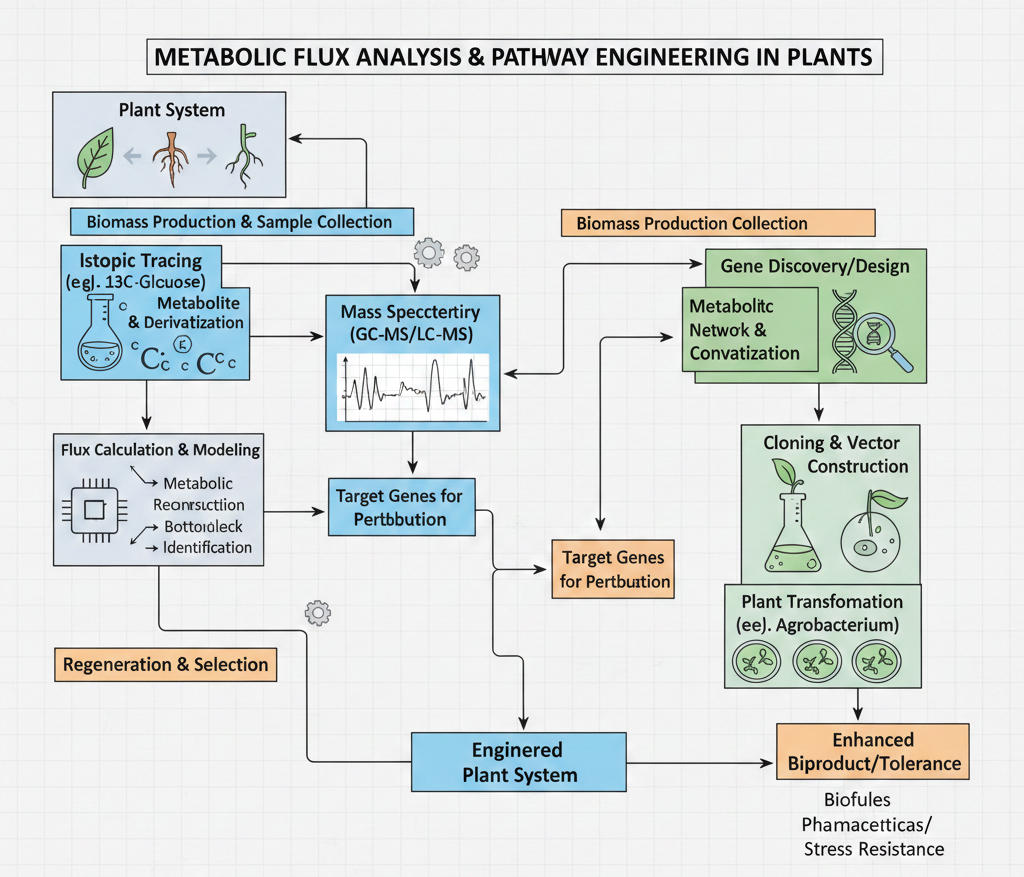
A schematic diagram for a metabolic flux analysis and pathway engineering experiment in plant system
Tools for metabolic flux determination
I. Experimental Tools
- Isotope Labeling Techniques
a. Radioisotopes
- Examples: ¹⁴C, ³²P, ³H, ³⁵S
- Application: Classic studies (e.g., Calvin cycle, glycolysis, TCA cycle) used radiolabeled substrates to track atom flow.
- Detection: Autoradiography, scintillation counting.
- Advantages: High sensitivity; excellent for low-abundance metabolites.
- Limitations: Hazardous; poor resolution for complex systems; limited dynamic range.
b. Stable Isotopes
- Examples: ¹³C, ²H, ¹⁵N, ¹⁸O
- Application: Used in ¹³C-Metabolic Flux Analysis (¹³C-MFA), Metabolic Flux Ratio Analysis, and Isotopically Non-Stationary MFA (INST-MFA).
- Detection: GC-MS, LC-MS, NMR.
- Advantages:
- Non-radioactive, safe for humans and environment.
- Enables detailed, quantitative, and compartment-specific flux analysis.
- Compatible with dynamic labeling experiments (e.g., pulse-chase).
- Analytical Platforms
a. Mass Spectrometry (MS)
- Used to detect isotopic enrichment in metabolites and determine isotopologue distributions.
- Often coupled with GC-MS or LC-MS/MS.
- Strengths: High sensitivity, throughput, and precision.
b. Nuclear Magnetic Resonance (NMR)
- Especially useful for ¹³C and ¹H-labeled metabolites.
- Allows non-destructive analysis and real-time monitoring.
- Strengths: Can quantify fluxes in intact cells and tissues (e.g., plant roots, perfused organs).
- Limitations: Lower sensitivity and resolution compared to MS.
II. Computational Tools and Models
- ¹³C-Metabolic Flux Analysis (¹³C-MFA)
- Uses isotopomer models and measured labeling patterns to compute intracellular fluxes.
- Requires:
- Known stoichiometric network.
- Accurate isotopologue distributions.
- Assumptions of metabolic and isotopic steady state.
Software tools:
- INCA (Isotopomer Network Compartmental Analysis)
- OpenFlux
- Metran
- FiatFlux
- Flux Balance Analysis (FBA)
- A constraint-based method that uses stoichiometry and optimization (usually of biomass or energy yield) to predict flux distributions.
- Assumes steady state and requires:
- A genome-scale metabolic model.
- Constraints (e.g., nutrient uptake, thermodynamics).
Software tools:
- COBRA Toolbox (MATLAB)
- RAVEN, CellNetAnalyzer, CarveMe
- Isotopically Non-Stationary MFA (INST-MFA)
- An advanced ¹³C-MFA method used when isotopic steady state cannot be reached, such as in fast-growing or photosynthetic organisms.
- Measures the time course of isotopic labeling after isotope introduction.
- Requires time-resolved metabolomics and ODE-based modeling.
- Dynamic Flux Analysis (dFBA)
- Models time-varying metabolic behavior under changing environmental or physiological conditions.
- Requires integration of fluxes over time, with time-course data for metabolites and biomass.
III. Emerging and Integrated Approaches
- Single-Cell MFA
- Using technologies like NanoSIMS, single-cell metabolomics, and microfluidic isotopic labeling, fluxes are being assessed at the individual cell level.
- Metabolic Control Analysis (MCA)
- Quantifies how sensitive fluxes are to changes in enzyme activity, metabolite levels, or transport rates.
- Helps identify control points in metabolic networks.
- Multi-Omics Integration
- Combining fluxomics with:
- Transcriptomics: gene expression profiles.
- Proteomics: enzyme abundance.
- Metabolomics: substrate/product levels.
- Enables systems-level understanding of metabolic regulation.
Pathway engineering in Plants
- Involves modifying plant metabolic pathways to enhance the production of desired compounds.
- Targets include primary metabolites (e.g., sugars, amino acids) and secondary metabolites (e.g., alkaloids, flavonoids).
- Approaches include gene overexpression, knockout, RNAi, CRISPR, and transgenic pathways.
- Synthetic biology enables the introduction of entirely novel pathways from microbes or other plants.
Pathway engineering in plants entails the intentional alteration of metabolic pathways to augment, divert, or initiate the biosynthesis of particular chemicals, aiming to boost plant performance, productivity, or the production of important biomolecules. This domain utilizes advancements in genetic engineering, synthetic biology, and systems biology to modify genes that encode enzymes, regulatory proteins, and transporters crucial to metabolic processes. Methods including the overexpression of essential enzymes, gene knockouts, RNA interference (RNAi), and CRISPR/Cas genome editing are frequently utilized to enhance flux through targeted pathways while reducing competing pathways. The integration of novel biosynthetic genes from different organisms facilitates the synthesis of non-native metabolites in plants, including medicines, nutraceuticals, and industrial precursors. Pathway engineering primarily aims to augment the synthesis of primary metabolites, including starch, lipids, and amino acids, to enhance nutritional value or biomass yield, alongside secondary metabolites like flavonoids, terpenoids, and alkaloids for stress resilience and medicinal uses. The efficacy of these alterations relies on a comprehensive understanding of pathway regulation, enzyme kinetics, and intracellular metabolite transport, frequently necessitating integration with metabolic flux analysis (MFA) and omics technologies. As research advances, pathway engineering offers significant potential for the creation of climate-resilient crops, sustainable bio factories, and innovative plant-based solutions to global issues.
Let us discuss some examples in plants where metabolic pathways were engineered to produce essential metabolites required for the growth and development of the human body. Carotenoids are one such class of essential molecules. They have multiple health benefits such as antioxidant, anti-inflammatory properties and also provide photo-protection against UV radiation. Most of the organisms including humans are unable to produce or synthesize carotenoids molecules, so there is a need for these pigments from the external sources such as plants and microalgae.
Vitamin A is essential for human health, and its lack leads to numerous vision-related disorders. Vitamin A deficiency is the primary cause of blindness in South Asian nations, and in severe instances, it may also result in morbidity and mortality among children [1]. Humans cannot synthesis vitamin A directly; therefore, the intake of diverse plant food sources such as oranges, spinach, carrots, and broccoli supply provitamin or vitamin precursors [2]. These dietary pro-vitamins facilitate the synthesis of vitamin A in the human body [3]. There are four types of plant provitamin A, also referred to as carotenoids. Alpha and beta carotene are present in greater quantities than gamma and beta-cryptoxanthin. The biosynthesis of plant carotenoids transpires in plastids, facilitated by enzymes encoded by nuclear genes. The advised daily dose of vitamin A is 700μg retinol activity equivalents for women and 900μg for males [4]. Nonetheless, minimal quantities of vitamin A are present in staple crops such as rice, millet, sorghum, potatoes, and other cereals prevalent in numerous nations. Genetic engineering techniques have effectively enhanced provitamin A production for the biofortification of many crops. The predominant technique to augment carotenoid production entails the overexpression of one or more genes that encode enzymes participating in this pathway. The incorporation of Phytoene desaturase from Pantoea ananatis and Phytoene synthase from Narcissus pseudonarcissus into the japonica rice cultivar results in the synthesis of biofortified golden rice, which contains 1.6 µg/g of carotenoid content [5]. Conversely, the incorporation of extremely efficient Phytoene synthase from Zea mays resulted in the accumulation of 37μg/g of carotenoids, transforming rice into Golden Rice 2 [6]. Likewise, canola, flax, tomato, potato, wheat, and soybean were genetically modified with individual or multiple carotenoid biosynthesis genes to yield the respective biofortified cultivars [7, 8, 9, 10, 11, 12] Additionally, another technique involves the silencing of genes expressing lycopene beta cyclase and lycopene epsilon cyclase to balance metabolic flow in favor of increased beta-carotene accumulation. This technique was employed in potato and wheat [13, 14, 15]. Concurrent overexpression of bacterial phytoene synthase (CrtB) and silencing of beta-carotene hydroxylase (HYDB) was demonstrated to enhance beta-carotene levels by up to 31-fold in wheat [15]. A third technique was implemented to aid in the stabilization of post-translational levels of genes associated with carotenoid production. This facilitated the sustained and ongoing accumulation of carotenoids in potatoes expressing the orange (OR) gene derived from cauliflower [16, 17, 18, 19]. Alongside the manipulation of carotenoid biosynthesis pathway genes, an alternative technique was implemented to guarantee the availability of isoprenoid precursors for carotenoid synthesis, circumventing the rate-limiting phase. This technique facilitated the development of Golden Rice 2 through the incorporation of the Arabidopsis ORANGE (AtOR) and AtDXS (1-deoxy-D-xylulose-5-phosphate synthase) genes, leading to markedly elevated carotenoid concentrations [20].
Applications
- Improving nutritional content (e.g., biofortified rice with higher iron or vitamin A).
- Enhancing biomass production for biofuels.
- Increasing resistance to stresses (drought, pests, salinity) via metabolic reprogramming.
- Producing high-value pharmaceuticals and industrial compounds in planta.
Challenges and Future Directions
- Complexity of plant metabolism and compartmentalization (e.g., plastids, mitochondria).
- Difficulty in precisely measuring fluxes in multicellular, heterogenous plant tissues.
- Need for better in vivo flux measurement tools and real-time monitoring systems.
- Integration of AI and machine learning for predictive metabolic engineering.
