Exploring RNA Secondary Structure Prediction using the Nussinov Algorithm
Procedure
Step 1.
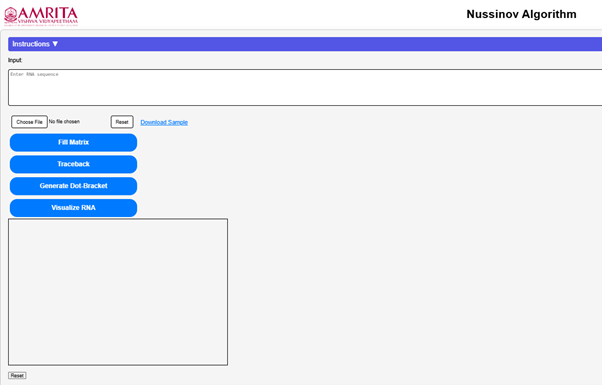
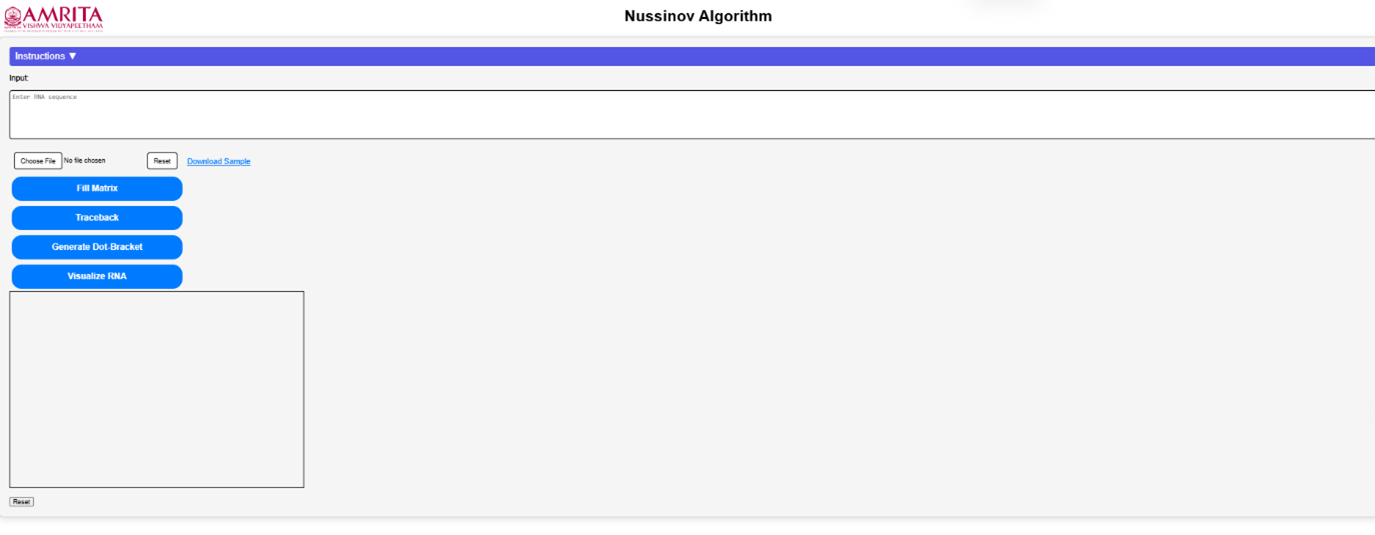
Open the simulator tab where user can find a user-friendly interface with different options to run the experiment.
Step 2.
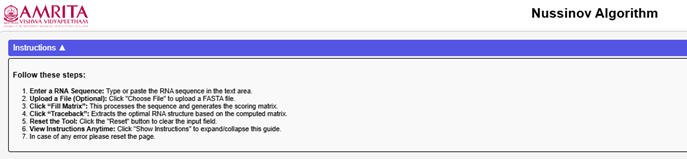
Click on the "Instruction" button. Read through the guidelines to understand how the simulator works before proceeding.
Step 3.
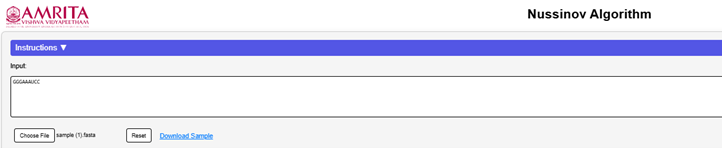
User can manually enter an RNA sequence into the input box. Alternatively, download a sample RNA file (. fasta format) from the page and upload it using the “Choose File” button.
Step 4.
Click on the “Fill the matrix” button to calculate the possible base pairs within RNA sequence
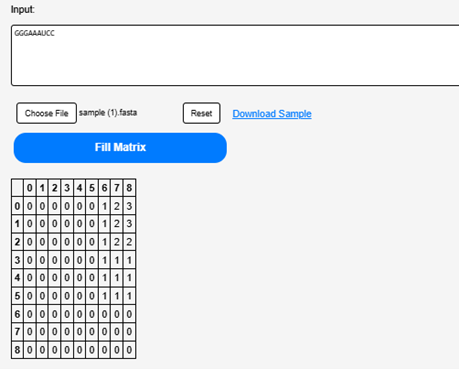
Step 5.
Click on the “Traceback” button to determine the possible secondary structures of uploaded RNA sequence.
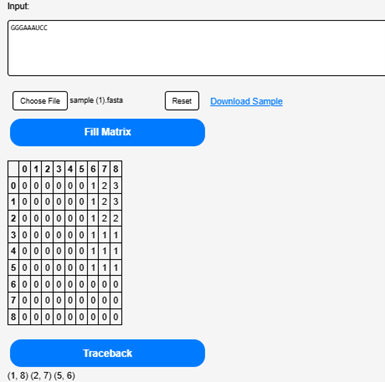
Step 6.
Click on the “Generate Dot-Bracket” to obtain a representation of the most stable RNA secondary structure.
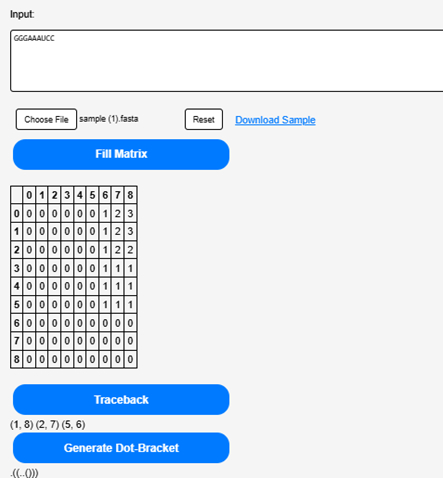
Step 7.
Click on the “Visualize RNA” to see a graphical representation of the most stable RNA structure.
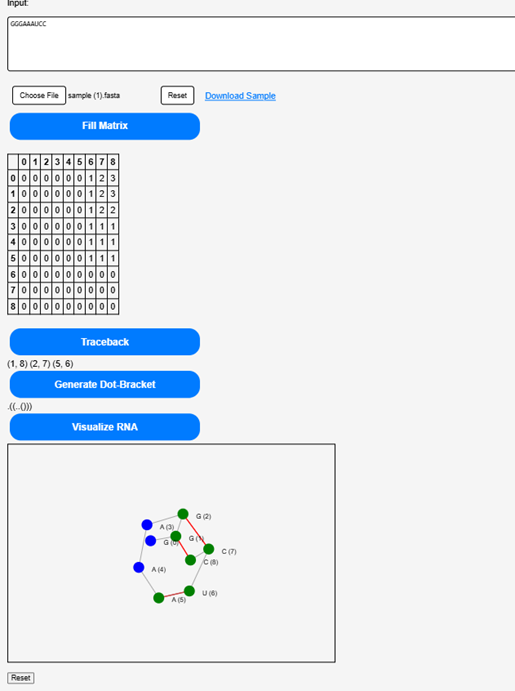
Step 8.
User can repeat the experiment, click on the “Reset” button to clear the current data and start again.